Notice: Trying to get property 'parent' of non-object in /home/nanonics/public_html/components/com_k2/router.php on line 502
Notice: Trying to get property 'alias' of non-object in /home/nanonics/public_html/components/com_k2/router.php on line 511
Super User
Schottky Diode I-V Characterization
 |
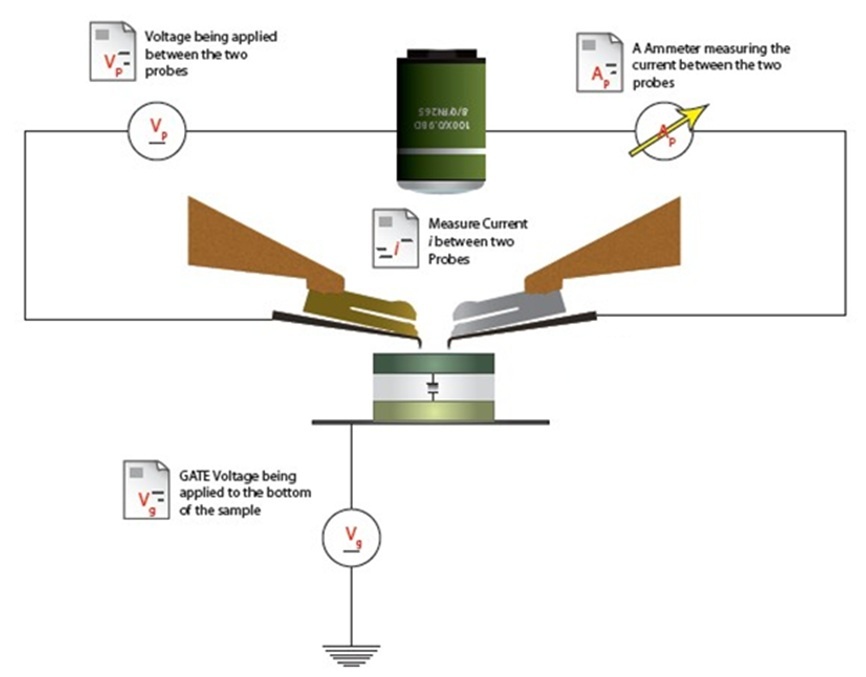 |
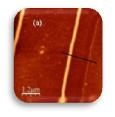
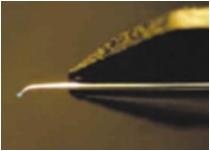
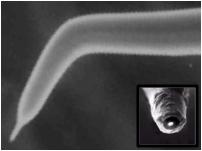
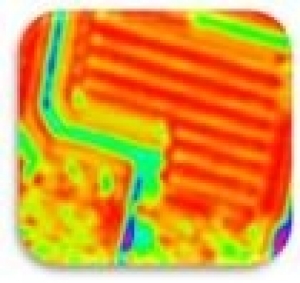
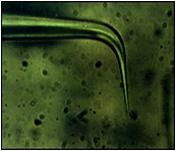
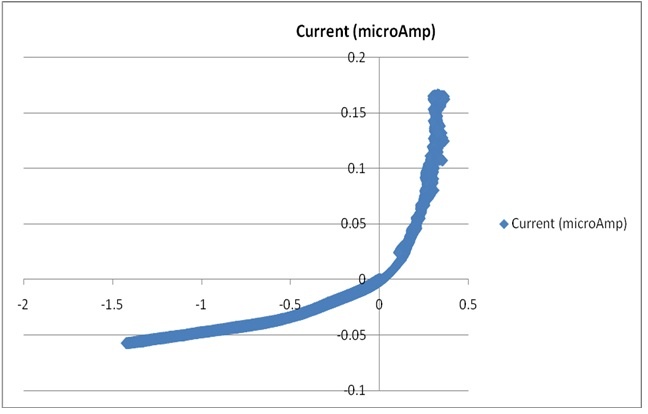
| Localized Schottky Diode I-V characterization: 20nm gold layer on 5nm Ti over Silicon substrate (Natural Oxide) | High sensitivity AFM Pt nanowires probes are used with the Nanonics MultiProbe system for electrical nano-charactarization of the inspected sample. One probe is used as a voltage source and the second probe is used to measure the current. The ability of these probes to provide both electrical and topographical information creates a special platform of accurate alignment at nanometric features such as nanotubes, nanoparticles and other molecular devices.
The electrical measurements and analysis are fully controlled through the SPM system. Hence, a comprehensive analysis of I-V nanometric spectroscopy correlated with topographical maps are provides. |
Electrophoretic lithography via electrical pulses for accurate and controlled chemical deposition
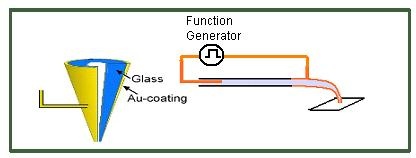 |
NanoInkJet printing through voltage controlled deposition |
Nanonics’ NanoInJet printing allows for writing with electric pulses across the tip of the nanopipette. This is obtained by applying a voltage between two electrodes located at the metal coating of the nanopipette tip and a Pt wire immersed in the liquid inside the nanopipette tube. This type of voltage allows for separation of molecules and the selective chemical writing of specific chemical species. This marries to great fields of chemical nanolithography and capillary zone electrophoresis. The voltage on the outside of the pipette can also be on the sample. Thus you have one electrode in the inside of the nanopipette and the other on the sample. For this the sample can be conducting or semiconducting but also a thin cover slip of glass coated with metal on the side opposite to what you are writing on.
| A) Positive or negative voltage pulses are provided between the metal coating at the nanopipette’s tip and the liquid at the inner area of the nanopipette | 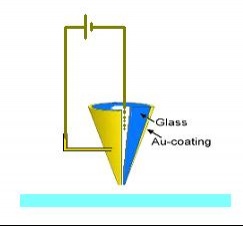 |
| B) Positive or negative voltage pulses are provided between the metal coating at the nanopipette’s tip and the sample | 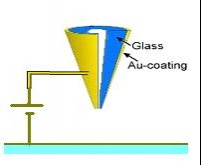 |
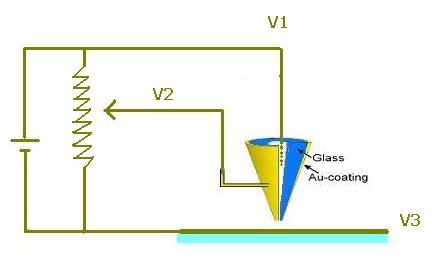
| C) Positive or negative voltage pulses are through three electrodes including nanopipette’s tip metal coating, sample and the liquid at the inner nanopipette area. |
 |
FPN Deposition of Different Inks on Variety of Samples
 |
A variety of inks can be injected into the nanopipettes through simple capillary actions without specific treatment. These include water, proteins, different buffers and metallic nanoparticles suspensions. In addition, a variety of surfaces can be used for lithography actions - including conducting, semi-conducting and non-conducting - such as glass, Si, Sio2. No special treatment is required for the sample. The flexible selection of inks and surfaces in FPN techniques allows for numerous advantages compared to other existing Nanolithography techniques such as NanoImprint, Dip Pen NanoLithography, Pulsed induced Lithography, and others.
|
|
|
|
|
|
|
Avidin on Aldehyde |
|||
|
|
|
|
|
|
|
|||
|
TiO2 on hydrophilic |
|||
|
|
|
|
|
|
|
|
||
Nanofountain Pen reviewed by Nature Materials
| 2003-08-14 | |||
Penning a protein patternNanoscale spots of proteins on a chip will allow high-throughput screening of protein expression and function. They can be written with a 'multi-colour' nanofountain pen. 14 August 2003 Philip Ball Patterns of proteins with nanoscale features can be written directly onto a surface using a 'nanofountain pen', a very narrow pipette attached to a scanning probe microscope. This technique should enable the preparation of 'protein chips' with a very high density of features that will function in an analogous manner to gene chips, allowing snapshots to be taken of a cell's active proteome. The method has been developed by Aaron Lewis of the Hebrew University of Jerusalem and co-workers1. It joins a scattering of other techniques for creating protein nanoarrays, but has significant practical advantages over these existing methods. DNA microarrays, in which strands of complementary DNA are attached in discrete, microscale patches to a solid substrate, have transformed genomic research by permitting a simultaneous analysis of thousands of genes to see which are activated at any moment. In principle, this enables the mapping of interactions and cooperation between genes associated with particular cell functions. But DNA arrays tell only part of the story. What really counts in the operation of a cell is the expression of proteins. There is not necessarily a direct relationship between gene expression and protein expression (some genes encode more than one kind of protein, for example). So molecular biologists are attempting to make protein microarrays in which tiny patches of surface-immobilized proteins bind to other proteins and reveal the expression patterns. The techniques used to make DNA microarrays can also work for proteins2. This produces spots 150–200 µm in diameter. But the smaller the spots, the greater the number of different proteins that can be assayed on a single chip. Chad Mirkin of Northwestern University in Illinois and co-workers have made protein nanoarrays using 'dip-pen nanolithography' (DPN), in which the tip of an atomic force microscope (AFM) is used as a kind of nanoquill3. It is dipped into some liquid 'ink' and then used to write nanoscale patterns, the molecules on the tip being transferred by capillary action onto a surface. To make this method work for proteins, Mirkin and colleagues have used two approaches. Proteins functionalized with thiol groups will stick directly to a gold substrate by a covalent bond. Alternatively, DPN was used to make template spots of other molecules, to which the proteins adhered3. Features as small as 100 nm across could be written this way. But the approach is not ideal for making arrays of many different proteins, which one way or another would require laborious multiple-dipping of the tip in as many different 'ink'. Lewis and colleagues have developed a different approach for delivering molecules to an AFM platform for nanoscale writing1. They attach a very fine pipette, with an aperture of just 100 nm or so, to an AFM cantilever arm and simply let the molecules flow out through the tip — they call it fountain-pen nanochemistry. Previously, the researchers demonstrated this method by depositing a chemical etchant to inscribe features in a metal film4. They have now adapted it for the creation of protein nanoarrays. They used the same protein system used previously by MacBeath and Schreiber in conventional microarray fabrication2: a yeast protein attached to an aldehyde-coated glass slide. The aldehyde reacts with the amine group of the protein's terminal amino acid to form a covalent linkage. In this way, Lewis and colleagues1 wrote lines of the protein about 500 nm wide, under ambient conditions — three orders of magnitude smaller than the features made by MacBeath and Schreiber. They then explored a different system: the green fluorescent protein (GFP) extracted from a Pacific jellyfish, which is widely used for molecular labelling in cell biology. The researchers wrote dots 250 nm across on a glass slide coated with bovine serum albumin, to which GFP binds. Even though this binding does not involve a covalent linkage, it is strong enough that the GFP features are not disturbed when they are repeatedly scanned for imaging with an AFM. Because of their fluorescence, they can also be clearly imaged with near-field scanning optical microscopy, which has subwavelength resolution. The crucial factor is that the 'fountain pen' can have different inks channelled into it automatically, simply by connecting it up to standard high-performance liquid chromatography instrumentation. This should make writing a multi-protein nanoarray much easier than by using DPN, and without the need for any complex pre-treatment of the substrate. In related work, David Klenerman of the University of Cambrdige in England and co-workers5 have used a nanopipette with an aperture of 90–130 nm to deposit proteins (antibodies), DNA and other biomolecules onto a surface with spot sizes of around 850 nm. The flow of molecules from the nanopipette was controlled by voltage pulses, which establish a very high electric-field gradient in the tip region and induce fluid flow by electro-osmosis. References
|
A NanoToolKit™
- A NanoToolKitTM of unique exposed tip with no Raman background probes
- All probes are MultiProbe friendly
|
AFM/NSOM
|
NanOptical Light Sensing: Near-Field Scanning Optical Microscopy - Illumination - Collection |
|
|
Nanopipettes
|
- NanoFountai Pen for liquids and gas delivery - NanoEvacuation - Ionic Conductance |
|
|
Electrical/Thermal
|
Glass insulated coaxial: - Nanoelectrical - Nanochemical probes |
|
|
Plasmonics
|
Tipped with single Au or Ag nanoparticle: - TERS - ANSOM (sSNOM) |
|
|
MultiProbe
|
MultiProbe: Imaging Nanomanipulation |
Uniquely capable with exposed tip for side wall imaging
- Permits ultra-sensitive imaging of side walls
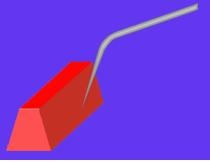 |
|
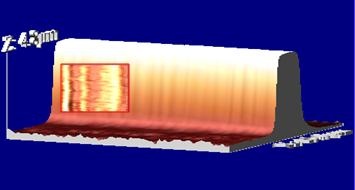 |
AFM 3D topographic presentation of a 4.3 microns step. Inset: AFM topographic image of the selected area on the step’s side wall. |
Elongated probe tip excellent for profiling
-
Comparative profiling probes with high aspects ratio for deep trenches
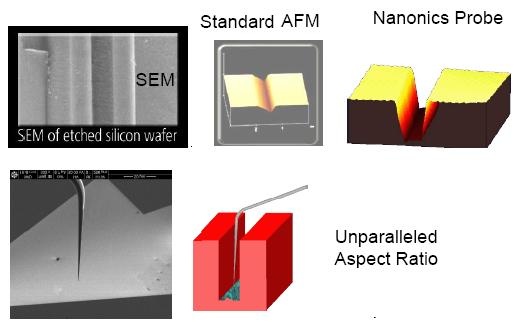
Optically friendly & completely non-interfering
-
Unlike standard silicon cantilevers such optically noninterfering probes provide a clear optical axis
-
The ultra-thin transparent glass tip with a high cantilever structure prevents any optical interfence even in Difference Interference Contrast (DIC) microscopy.
| Conventional AFM Limitations | Nanonics’ non-interfering glass probe |
 |
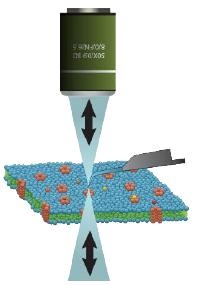 |
| Si probe’s cantilevers cause shadowing and Raman interference | Glass probes are transparent with exposed tip geometry. |
Transparent probes with exposed tips & no Raman background
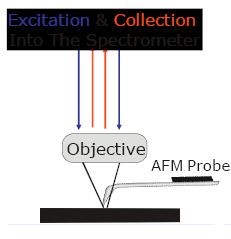 |
 |
| Transparent and non interfering cantilevered glass probe are idea for SPM/Raman/TERS applications |
|